Critical care - and the medical sector generally - is lagging behind other industries when it comes to implementing the latest technology into its practices
New technologies in the critical care sector have the potential to benefit patients and doctors greatly (Credit: Wikimedia)
In the past few decades, we have witnessed a particularly rapid period of technological advancement in the area of critical care, with data and the need to interpret it now an integral part of the field. Practical Patient Care writer Emma Green explores these developments in technology and considers the implications for optimising patient care.
It is clear that 21st-century healthcare requires intensive use of technology to acquire, analyse, manage and disseminate data.
Nowhere is this more critical than in the intensive care unit.
While there have been major improvements, the medical industry, for the most part, has not yet fully incorporated many of the advances in computer science, biomedical engineering, signal processing and mathematics that many other industries have embraced.
Despite the growth of critical care, the basic approach of data collection and management has remained largely stagnant over the past 40 years.
Large volumes of information are collected from disparate sources and reviewed retrospectively.
This is highly challenging in itself as providers are required to navigate through a jungle of monitors, screens, software and often paper.
Data from patient monitors and medical devices, although sometimes visible at bedside, is difficult to acquire and store in digital format.
Currently, there is limited medical device interoperability and integration with the electronic medical record (EMR) remains incomplete and cumbersome.
In addition to these limitations, standard analytical approaches provide little insight into a patient’s pathophysiologic state, which is imperative to understand the dynamics of critical illness.
In order to optimise care in this context, healthcare professionals need precisely time-stamped data, integrated with clinical context and processed with a range of analytical tools.
These demands are often beyond the capability of typical commercial monitoring systems.
A comprehensive understanding from advanced data analytics can aid physicians in making timely and informed decisions, and improving patient outcomes.
Ultimately, an integrated critical care informatics architecture will be required, which includes acquisition, synchronisation, integration and storage of all relevant patient information into a single, searchable database, as well as the ability to gain practical insights from this data.
A brief history of computers and critical care technology in ICUs
The Electronic Numerical Integrator and Computer (ENIAC) was the first general-purpose computer and was introduced in 1946.
It was developed to calculate missile trajectories for the US Army and was the size of a room, weighing 27t.
A few years later, commercially available computers hit the market, but due to their high cost their use was limited to large corporations to manage their accounting.
In the 1960s, hospitals began to develop EMR systems including the Problem Oriented Medical Record (POMR) at the University of Vermont, Health Evaluation through Logical Processing (HELP) at the University of Utah, The Medical Record (TMR) at Duke University and the Computer Stored Ambulatory Record (COSTAR) at Harvard.
Although these early systems were capable of processing medical information, they were rarely connected to the real-time data-intense environment of the ICU (intensive care unit) and thus has limited applicability in this context.
The first computer to be introduced into critical care was in 1966 and was used to collect vital signs from the bedside monitor automatically.
Through connecting this device, it was possible to obtain arterial and venous pressure, heart rate, temperature and urinary output.
This had been previously attempted in 1934 with a mechanical contraption but was not entirely successful.

Basic analytical tools, such as trend analysis, were later added to the automated data collection system to improve functionality.
Other early applications of computers in healthcare included clinical decision support systems to assist in the diagnosis of haematologic disorders, tools for respiratory monitoring and automation of blood transfusion after cardiac surgery.
For example, the computer-based Clinical Assessment, Research and Education System (CARE) was a clinical decision support system designed to help with the treatment of critically ill surgical patients.
This continuously monitored physiologic and metabolic markers and managed data about fluid and electrolytes as well as cardiac and respiratory functioning.
In the 1980s, automatic collection of heart rate and blood pressure became increasingly advanced with data being presented in graphical displays instead of bedside flow sheets.
The architecture also improved from the locally contained model to the client/server model in which a workstation in the ICU interacted with a central computer housing patient data via a local area network.
Links to the hospital EMR systems were also being made, such as computers systems that handled admissions, discharges and transfers so that patient demographic data could be readily accessed by healthcare professionals.
Physician and nursing notes were later able to be entered electronically into problem-oriented medical records.
Computers were also being introduced into the operating room, with computerised anaesthesia records allowing for more reliable collection, storage and presentation of data during the perioperative period, in addition to providing basic record-keeping functions.
However, data from medical devices were rarely integrated with the other physiological information.
The current state of clinical information systems in critical care
Today, there are a number of commercially available clinical information systems for the ICU.
These have continued to evolve over the years, with various acquisitions resulting in the creation of broad end-to-end platforms.
While these represent a significant improvement compared with past technology, there are several existing limitations.
Currently these systems are restricted in terms of functionality and the acquisition of high-resolution physiologic data.
This is due to a trade-off between the memory requirements of capturing high-resolution physiological data versus capturing data snapshots that may be sufficient for some clinical decisions.
Standards have yet to be set about where that balance lies.
Despite the increasing amount of information collected, visual displays in the ICU have remained largely unchanged for the past several decades.
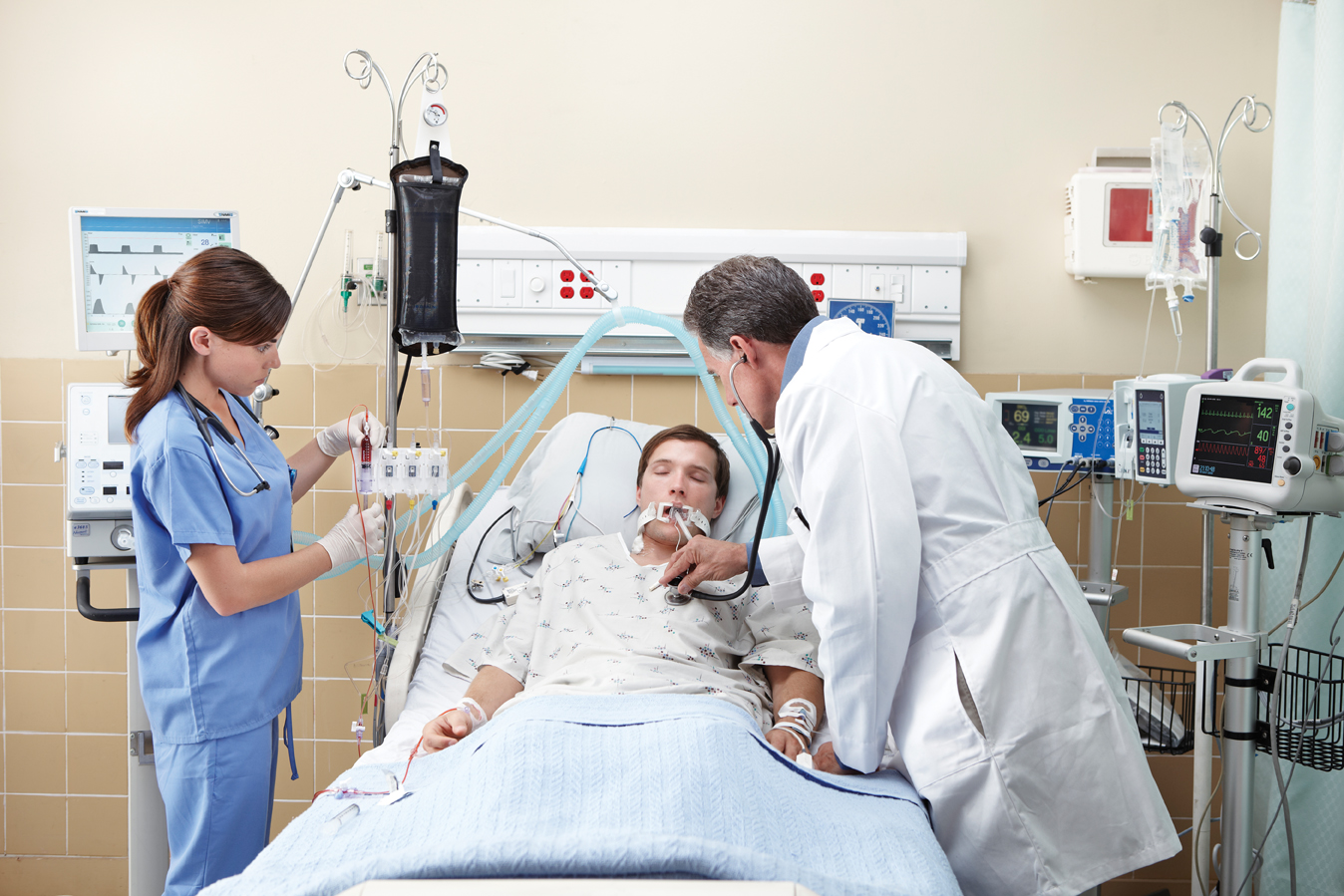
Clinicians can be confronted with more than 200 variables when caring for critically ill patients, yet most people cannot judge the degree of relatedness between more than two, which can contribute to medical errors.
In order to prevent this from occurring, graphical displays must be mindfully designed by applying a human systems integration approach.
It is important to understand not only how information should be optimally presented to promote a better understanding of the patient’s pathophysiologic state and support decision-making, but also to facilitate collaboration and optimal work-flow among the whole healthcare team.
The promise of critical care informatics lies in the potential to use these advanced analytical techniques on high-resolution multimodal physiological data to obtain more knowledge of the complex relationships between physiological parameters, improve the ability to predict future events and thus provide targets for individualised treatment in real time.
Future systems will go beyond simply reporting streams of raw data, but will synthesise it to generate hypotheses that best explain the observed data, providing situational awareness to the clinician.
Medical device interoperability and data integration
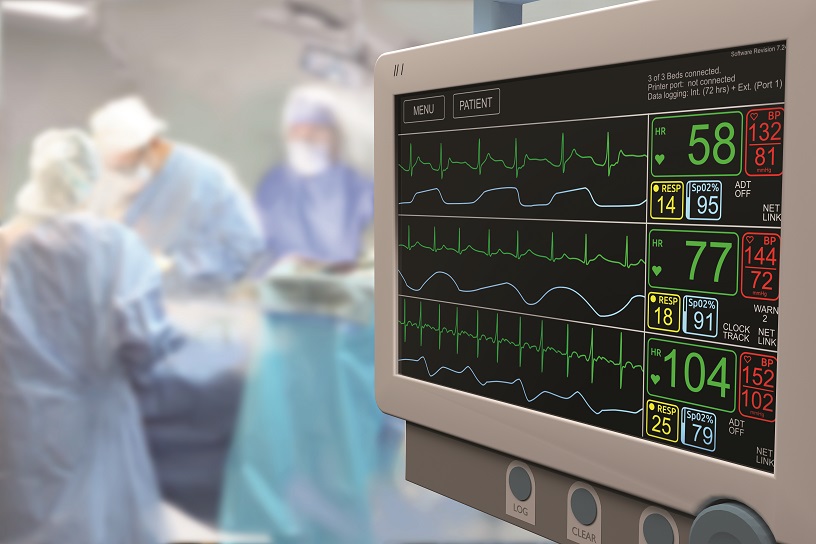
Central to the growth of critical care has been the increase in monitoring technology and stand-alone medical devices.
A wealth of information is generated by reflecting dynamic and complex physiology, which can only be understood through integrating data with the clinical context.
However, the vast majority of these variables are generated from individual devices that are not readily compatible with each other.
Some connect directly into the bedside monitor but many only do this partially, if at all, which means that not all data is captured electronically.
The lack of interoperability is one of the most significant limitations not only within critical care but within healthcare more generally.
This is in stark contrast with the ‘plug and play’ capabilities of consumer electronics.
Many groups are tackling the problem of interoperability on their own by developing the hardware and software interfaces that facilitate device connectivity.
Connecting with analogue data ports demands appropriate hardware interfaces, analogue-to-digital (A/D) converters, and filters to eliminate aliasing due to a mismatch between sampling rate and the frequency content of the signal being acquired.
It also requires that the data be properly scaled to the voltage range of the A/D converter (microvolts to millivolts) to maximise the resolution.
Although such approaches provide the opportunity to individually interface with a variety of devices in the ICU, a system that provides comprehensive, cross-manufacturer medical device integration for the care of a single critically ill patient at the bedside is not yet available.
When data is being acquired from different devices, each with its own internal clock, the time stamps of data acquired simultaneously can all be different.
In order to align these, time synchronisation of the information is critical.
Furthermore, even when acquiring data from a single patient monitor, time drifting from natural degradation, daylight savings time or incorrect adjustments made by the clinical staff need to be rectified.
Without a universal clock ensuring that all the values are in sync, interpreting the information is highly challenging, if not impossible.
Data acquisition and integration systems
Commercial off-the-shelf products do not typically support high-resolution physiologic data acquisition, archiving, or annotation with bedside observations for clinical applications.
This is largely because such systems have been developed in academic settings mainly for clinical research.
As they are not open source, most of these are not readily available, which has resulted in substantial duplication of effort in software development for acquiring and archiving physiological data.
There has been considerable effort to address this issue, ranging from developing and testing of new mathematical and analytical tools, to hardware and software solutions for patient data acquisition, archiving and visualisation.
Some have also focused on multimodal data collection linked with clinical annotation.
While there have been significant improvements in intensive care monitoring, there remains a lot of untapped potential to capitalise on recent advances in computer science, biomedical engineering, signal processing and mathematics.
Acquiring, synchronising, integrating and analysing patient data remains highly challenging due to the lack of sufficient computational power and a lack of specialised software, incompatibility between monitoring equipment and limited data storage within current hospital systems.
As a result of recent developments in technology, all of these technical problems are now surmountable.
Today, we are fortunate to be living within a data-intensive science era in which there is a wealth of information available to generate insights that can be used to optimise the speed and accuracy of clinical decision-making, improving the lives of patients.
This article first appeared in Issue 23, 2019 of Practical Patient Care. The full publication can be viewed online here.
